Published online 2005 Jun 17. doi: 10.1251/bpo108
Quick and Dirty Screening for Cloned Inserts. However, there is a quick & dirty screening method that is even faster than colony PCR. Called colony cracking, it involves picking a colony straight into lysis buffer, running the extract straight onto an agarose gel, and then identifying positive clones based the electrophoretic mobility difference. As for the PCR; you should try using one primer that binds to the vector and one primer that binds to the insert. A PCR product will tell you insert in in your plasmid (at least the end which was sequenced). You may then want to check the other end of the fragment.
PMID: 16136227
Abstract
We have developed a simple and effective method (Lig-PCR) for monitoring ligation reactions using PCR and primers that are common to many cloning vectors. Ligation mixtures can directly be used as templates and the results can be analyzed by conventional gel electrophoresis. The PCR products are representative of the recombinant molecules created during ligation and the corresponding transformants. Orientation of inserts can also be determined using an internal primer. The usefulness of this method has been demonstrated using ligation mixtures of two cDNA’s derived from the salivary glands of Aedes aegypti mosquitoes. The method described here is sensitive and easy to perform compared to currently available methods.
Keywords: Ligation, Nucleic Acids, Polymerase Chain Reaction
Introduction
Ligation is a vital process in the normal replication, repair and recombination of nucleic acids (, ). In vitro, it is an important tool for editing DNA and is frequently used for cloning of products generated by restriction enzyme digestion, PCR, RT-PCR and also for construction of genomic (), cDNA () or phage display libraries (). Some methods like Ligase Chain Reaction (), Amplified Fragment Length Polymorphism () and detection of mononucleotide repeat sequences (8) depend on nucleic acid ligations, while synthetic circular RNA molecules have also been made in vitro using RNA ligation (). Being an essential component of many molecular biology procedures and having an increasing range of application, the success of ligation can become a critical parameter. Ligation efficiency becomes important during the generation of complex libraries (10-12) and with some of the above applications which involve ligation as an intermediate step. Hence, being able to evaluate products of a ligation reaction can be very helpful in planning subsequent steps. Creation of recombinants by ligation and their subsequent transformation is a relatively time-consuming process, with hardly any intermediate screening points. A simple method would be to check the ligation products by gel electrophoresis, but sacrificing a large portion of the ligation mixture (typically 10-30%) for conventional gel electrophoresis seems impractical. One of the methods previously described for assaying ligation reactions involved the labeling of ligation products using 32P, followed by denaturing gel electrophoresis and autoradiography (, ). In addition to not being sensitive, this method is time-consuming, discontinuous and involves the use of radioactivity. Other methods involve the use of capillary electrophoresis (), real-time monitoring of ligations using molecular beacons () and an enzymatic bioluminescent assay (). These methods have their own advantages, such as sensitivity (real-time monitoring using molecular beacons) or using very little sample (capillary electrophoresis). But they are specialized and require the availability of special reagents and equipment, to which many labs may not have access. Here, we describe a simple and effective method (which we have termed Lig-PCR) to analyze or monitor ligation reactions using PCR () and M13 based primers that are common to most cloning vectors (). Recombinant sequences can be directly amplified from very small quantities (0.5-2% or even lower, depending on the amount of plasmid molecules and number of PCR cycles) of ligation mixtures, without the need for intermediate purification steps. The results can be visualized on a standard agarose gel and completed in the time that is normally available between ligation and transformation (2-3 hours). An additional feature of this method is the capacity of determining the orientation of inserts, using an internal primer based on the insert sequence.
Materials and Methods
Generation of inserts for TA ligation (ligation of PCR products generated by Taq DNA polymerase to T overhang vectors) and cohesive ligation
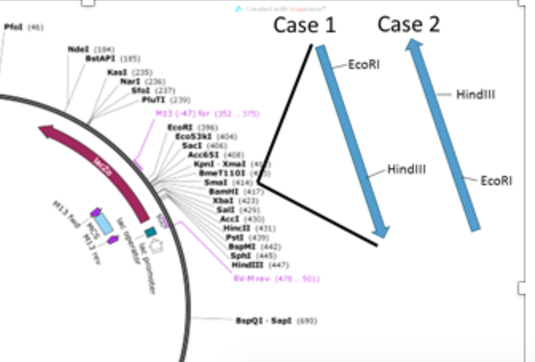
We used two PCR products (60S-L37 and 40S-L17) derived from an Aedes aegypti mosquito salivary gland cDNA library as inserts for the ligation experiments described here. These PCR products correspond to the 60S ribosomal protein L37 cDNA (60S-L37; GenBank Acc. No. AY552059) and the 40S ribosomal protein L17 cDNA (40S-L17; GenBank Acc. No. AY927787) from this mosquito. The original cDNA library was constructed in a λ TriplEx2 vector (BD Bioscience/Clontech, Palo Alto, CA, USA) and the 60S-L37 and 40S-L17P inserts were generated as PCR products using vector based primers 5’ λ TriplEx2 sequencing primer and λ TriplEx3’ LD-insert screening amplimer (Fig. (Fig.1A1A and Table Table1).1). The cDNA synthesis procedure uses a primer CDS III/3’ PCR primer [ATTCTAGAGGCCGAGGCGGCCGACATG-d(T)30N-N], which gets incorporated at the 3’ end of all the cDNA’s. A primer designed from the above sequence is used as a common insert specific primer (Fig. (Fig.1).1). The PCR products were purified using the AMPure® solid-phase paramagnetic beads (Agencourt Biosciences, Beverly, MA, USA) and used for TA cloning into pCR 2.1 TA vector (Invitrogen, Carlsbad, CA, USA). For cohesive ligation, plasmid vector pCR 2.1 (0.5 mg) and inserts 60S-RP and 40S-RP (1 mg each) were double digested separately with 5 units each of restriction enzymes EcoR I and Hind III (Invitrogen, Carlsbad, CA, USA), purified using phenol-chloroform extraction and ethanol precipitation (20) and then subsequently used for ligation.
Strategy used for generating inserts for ligation. A. Generating inserts for 60S-L37 and 40S-L17 using PCR from Aedes salivary gland cDNA clones for TA and cohesive end cloning. CDS-III 3’ primer used as insert specific primer. B. pCR 2.1 plasmid vector (Invitrogen, Carlsbad, CA, USA) used for TA and cohesive end cloning (Map not to scale. Complete vector sequence and map can be found at www.invitrogen.com).
Table 1
Primers used for cloning and analysis of ligation mixtures.
| Primer | Sequence (5'-3') |
| 5' λ TriplEx2 sequencing primer | TCCGA GATCTGGACGAGC |
| λ TriplEx 3' LD-amplimer | ATACGACTCACTATAGGGC GAATTGGC |
| M13-R (Reverse) | CAGGAAACAGCTATGAC |
| M13-F (Forward) (-20) | GTAAAACGACGGCCAG |
| CDS III/3' primer | CCGAGGCGGCCGACATG |
Ligation
Twenty five nanograms each of 60S-L37 and 40S-L17 purified PCR products (referred from now as insert) were ligated to 25 ng of either TA or EcoR I and Hind III digested pCR 2.1 plasmid, using 2 units of T4 DNA ligase (Invitrogen, Carlsbad, CA, USA). The vector to insert ratio was about 1:6 in a final volume of 10 mls and the reaction was carried out at 14°C for 14-15 hrs in a bench top cooler (Tropicooler 260014, Boekel Scientific, Feasterville, PA, USA). The liagtion mixtures were used either directly for PCR (referred from now as Lig-PCR) or for transformation of chemically competent One Shot® Top10 E.coli cells (Invitrogen, Carlsbad, CA, USA).
PCR amplification from the ligation mixture (Lig-PCR)
The ligation mixtures were diluted 1:10 times with milli-Q water and 1.0 ml of each ligation mixture (1%) was used to PCR amplify recombinant sequences (vector and insert) using M13-reverse and M13-forward (-20) plasmid specific primes (Fig. (Fig.1B1B and Table Table1).1). The PCR was carried out in a GeneAmp® PCR system 9700 (Applied Biosystems, Foster City, CA, USA) and the reaction consisted of 0.25 mM of each primer, 180 mM of each dNTP and 0.5 units of Taq DNA polymerase (Eppendorf/Brinkmann, Westbury, NY, USA) in a final volume of 20 ml. The cycling conditions were an initial denaturation at 95°C for 1 min., followed by 35 cycles of 95°C for 30 sec., 60°C for 20 sec., and 72°C for 1 min. The amplification products (referred as Lig-PCR products) were separated on a 1.5% Agarose/EtBr gel and visualized using a ChemiImager 4400 Imaging system (Alpha Innotech Corporation, San Leandro, CA, USA). The PCR products were quantitated as mean grey values of each band using ImageJ 1.33 software (http://rsb.info.nih.gov/ij/) and expressed as percentage of recombinant or vector derived products for each ligation mixture.
Transformation
Two microliters of undiluted ligation mixture were added to 25 mls of chemically competent One Shot® Top10 E.coli cells (Invitrogen, Carlsbad, CA, USA) and mixed gently. After 30 minutes on ice, the cells were heat shocked at 42±C for 30 sec. without shaking and immediately transferred on to ice. To the transformed cells, 125 μls of S.O.C medium (2.0% Tryptone, 0.5% Yeast extract, 10mM NaCl, 2.5mM KCl, 10mM MgCl2, 10mM MgSO4, 20mM glucose) was added and incubated at 37±C for 1 hour, with shaking at 225 rpm. The transformation mixture was spread equally on to two LB agar plates with Kanamycin (50 μg/ml) and X-gal (0.5 mM). Plates were incubated overnight at 37±C and blue and white colonies were counted the next day. Distribution of recombinant (White) and plasmid specific (Blue) colonies for each ligation mixture is represented as percentage.
Results and Discussion
In Fig. Fig.1A1A & B, the method that was followed to generate the insert for ligation reactions is outlined. The 60S-L37 and 40S-L17 PCR products are around 660 and 600 bp respectively. The Taq DNA polymerase generated PCR products (with A overhang) were ligated directly to pCR 2.1 TA vector; while for cohesive ligation, the PCR products were digested with Eco RI and Hind III restriction enzymes, resulting in a shorter insert size by about 124 bp each (Fig. (Fig.1A).1A). Eco RI and Hind III digestion of plasmid pCR 2.1 results in the deletion of a short 65 bp fragment (Fig. (Fig.1B).1B). Figure Figure22 shows the Lig-PCR products obtained by direct PCR amplification of TA and cohesive ligation reactions using M13 forward (-20) and M13-reverse primers (Fig. (Fig.1).1). The sizes of PCR products are listed below the gel profile, with the lower bands of 200 bp corresponding to the product obtained from the plasmid vector without any insert. Clear differences are visible between the different types of recombinant molecules created in the two reactions, based on the corresponding insert size. The distribution of recombinants compared to the re-ligated plasmid (described here as percentage of PCR products) was higher for cohesive ligations (Fig. (Fig.2,2, Lane 4 & 5), than for TA ligations (Fig. (Fig.2,2, Lane 2 & 3). TA ligations for both 60S-L37 and 40S-L17 show a high percentage products derived from re-ligated plasmid vector (Fig. (Fig.2,2, band 2), similar in size to the ones obtained from control, undigested plasmids (Fig. (Fig.2,2, Lane 1). The 60 bp size difference between the 60S-L37 and 40S-L17 insets (Fig. (Fig.1A;1A; Fig. Fig.2,2, Lane 6 & 7) is clearly visible in the corresponding Lig-PCR products in both TA and cohesive ligation mixtures (Fig. (Fig.2,2, Lanes 2, 3 & 4, 5, respectively). As depicted in Figure Figure1,1, the recombinants generated from cohesive ligations would be shorter by about 190 bases than their TA ligation counterparts, which can be seen in the corresponding PCR products generated using this method (Fig. (Fig.2,2, Lane 2, 4 & 3, 5 respectively).
PCR products from ligation mixtures using vector specific primers M13-R & M-13F. Electrophoresis on 1.2 % agarose gel and staining with Ethidium Bromide. A: Band 1 derived from recombinants and Band 2 from relegated pCR 2.1 vector. Lane 2 & 3 has two types of recombinant specific products and both were used for calculation band intensity/amount. B: PCR product amounts calculated as corresponding band intensity (mean grey value) using ImageJ 1.33 software (http://rsb.info.nih.gov/ij).
This is consistent with high ligation efficiencies expected for cohesive-end ligation, relative to TA ligations. These results also demonstrate the simplicity and usefulness of this method to monitor ligation mixtures using PCR. Since only a small portion of the reaction is required, without any intermediate purification step, the time required for this analysis usually fits between typical ligation and transformation procedures. This method was also repeated using purified ligation mixtures and the results were comparable (data not shown). This ability to screen ligation mixtures becomes important in planning subsequent steps, thereby saving time, reagents and effort.
To demonstrate the ability of Lig-PCR method in predicting the results of the transformantion, a portion of the above ligation mixtures were transformed into chemically competent E.coli cells and the resulting colonies were counted. Figure Figure33 shows the relative amounts of recombinant and vector specific colonies obtained from TA and cohesive ligations for 60S-L37 and 40S-L17, expressed as percentage. Cohesive ligation mixtures resulted in around 86% and 77% of recombinants for 60S-L37 and 40S-L17, respectively; while for TA ligation mixtures, this number was around 52% and 43%. This ratio is similar to the corresponding Lig-PCR band intensities (Chart B, Fig. Fig.2).2). Same type of correlation existed for transformants resulting from re-ligated plasmid vectors. The nature and amounts of Lig-PCR products can be used as an indication for the expected transformants from ligation mixtures, further supporting the usefulness of this method.
Distribution of transformants and Lig-PCR products from ligation reactions. Graph showing the relative percentages of transformants for each ligation mixtures. Colonies counted after blue-white screening on X-gal media. Also, confirmed by PCR (data not shown). Error bars: < 5%. Chart A: Recombinant and vector specific transformants expressed as percentage. Chart B: Recombinant and vector specific Lig-PCR products expressed as percentage (shown for comparison). The charts represent data from three independent experiments. TA & Coh-60S & 40S: Transformants obtained from TA and Cohesive ligation samples for 60S-L37 and 40S-L17. Refer to Figure 2 for description of Ligation samples.
We have also used the Lig-PCR method to determine the relative orientation of inserts in recombinants generated from the above ligation reactions. An internal primer on 60S-L37 and 40S-L17 cDNA insert, CDS III/3’ primer (Fig. (Fig.1)1) was used in combination with either M13-forward (-20) or M13-reverse primer on pCR 2.1 vector for PCR amplification of TA and cohesive ligation mixtures. Figure Figure44 shows the profile of products obtained (Lanes 1-8) in comparison to controls (Lanes 9-12), where no insert specific primer was used. The gel profile indicates that in the case of both TA and cohesive ligations for 60S-L37 and 40S-L17, relatively more inserts were in M13-Forward (-20)-CDS III/ 3’ primer orientation (Lane 1, 3, 5, 7) than in the M13-Reverse-CDS III/ 3’ primer orientation (Lane 2, 4, 6, 8). As determined by band intensity values, the percentages of recombinants in the M13-Forward primer orientation were around 88% and 68% for TA ligated 60S-L37 and 40S-L17; while in case of cohesive ligations, they were around 73% and 71%, respectively. Interestingly, inserts in both types of ligation mixtures showed similar types of preferential orientations, which was only expected for Eco RI-Hind III directional cloning (Fig. (Fig.1).1). The Lig-PCR products in Lanes 4 and 8 of Figure Figure44 are from single restriction enzyme digests, where the insert can orient in both directions. The higher bands seen in many lanes can result from concatemer molecules.
Determining insert orientations using Lig-PCR method. All PCR reactions (except controls) were carried out with the insert specific primer CDS III-3’ (Table 1) and either M13-F or M13-R. TA: TA ligation; Cohesive: Cohesive (EcoR I-Hind III) ligation. F: M13- forward primer; R: M13-reverse primer.
In summary, the method described here complements other methods that have been developed to monitor nucleic acid ligation reactions (-). More importantly, the Lig-PCR is a simple and effective method for analyzing ligation mixtures without the need for any intermediate purification steps, special reagent, equipment or radioactivity. To our knowledge, this is the only method that can be used for determining insert orientations directly from ligation mixtures, as has been demonstrated in the results (Fig. (Fig.4).4). This becomes useful during non-directional ligation in cloning and expression vectors. We have used around 1% of the ligation mixture for our analysis, a higher amount compared to about 0.02% mentioned for capillary electrophoresis based analysis (), but using an amount lower than 1% is also possible. The minimum amount of sample required for Lig-PCR analysis has to be standardized and will depend on the size and amount of plasmid vectors being used and the complexity of the ligation mixture. Being a PCR based method and resulting in amplification of the initial template, the Lig-PCR method is preferable over other sensitive but complicated methods (-). The average time required to carry out the Lig-PCR method would be about 2-3 hours and can depend on the length and number of PCR cycles, followed by gel electrophoresis. This method can be customized for most ligation mixtures and it is possible to reduce the total time required to carry out this procedure. Any set of vector based primers, which covers a cloning site, can be used and the amplification cycles can be adjusted according to the template and primers. Usually, these parameters are already standardized for regular PCR analysis. A Lig-PCR product band from agarose gels can be directly eluted, purified and used as a template for cycle sequencing. This can be another application in cases where the insert sequence or suitability of amino acid reading frame needs to be determined, without going through the normal transformation, colony screening and plasmid isolation procedures.
Potential difficulties and challenges that can be encountered during designing and performing this method may include designing primers for plasmids that do not have primer binding sites on either sides of cloning sites. Most commercially available plasmid cloning vectors have such sites for M13, T3 and T7 or other universal primers. Before performing the Lig-PCR, primer annealing and extension conditions will have to be standardized using regular PCR. Quantitating the PCR products on gel will require a gel documentation system and an image analysis or software program. We have used ImageJ 1.33 for this purpose, which can be freely downloaded from http://rsb.info.nih.gov/ij/. For determination of insert orientations, proper controls should be used to ensure that the PCR products are derived only from recombinant molecules in the ligation mixture. Any laboratory having access to basic molecular biology infrastructure can perform this analysis on a regular basis. We have ourselves found it very useful for planning cloning strategies and troubleshooting unexpected results.
Acknowledgments
This research was supported in part by U.S. Army Medical Research and Material Command award DAMD 17-03-1-0075 to S.K.W. The authors have no conflict of interest to declare related to this publication.
Appendix
Protocols
Equipment
- Thermal cycler: GeneAmp® PCR system 9700 (Applied Biosystems, Foster City, CA, USA)
- Bench top cooler: Tropicooler 260014 (Boekel Scientific, Feasterville, PA, USA).
- Temperature controlled water bath: Isotemp 105 (Fisher Scientific, Pittsburgh, PA, USA)
- Incubator Shaker: Classic C24 Series (New Brunswick Scientific, Edison, NJ, USA)
- Gel Electrophoresis Apparatus: MGU 202T & MGU 602T (C.B.S. Scientific, Del Mar, CA, USA)
- Gel Imaging System: ChemiImager 4400 Imaging system (Alpha Innotech Corporation, San Leandro, CA, USA)
Reagents
- Plasmid cloning vector: pCR 2.1 plasmid (Invitrogen, Carlsbad, CA, USA)
- Restriction enzymes: EcoR I, Hind III
- DNA ligase: T4 DNA ligase
- Competent cells: One Shot® Top10 E.coli chemically competent cells (Invitrogen, Carlsbad, CA, USA)
- Milli-Q water
- Plasmid specific primers: M13-forward and M13-reverse primers
- PCR reagents: 10X reaction buffer, dNTP mix (2.5 mM each) and Taq DNA polymerase Eppendorf/Brinkmann, Westbury, NY, USA)
- Agarose: SeaKem LE agarose (BioWhittaker Molecular Applications, Rockland, ME, USA)
- Ethidium Bromide.
- S.O.C medium (premixed, Invitrogen, Carlsbad, CA, USA)
- Bacterial growth media: Luria-Bertani medium (premixed dry powder, Fisher Biotech, Pittsburgh, PA, USA
- Antibiotic: Kanamycin (50mg.ml stock)
- Blue-White screening for recombinants: Using X-Gal (0.5 mM final) in the growth medium
Method
- Perform ligation reactions as usual. Use about 25 ng of plasmid vector per 10 μl reaction, in order to have sufficient templates for the subsequent PCR amplification.
- Dilute the ligation mixture 1:10 times with milli-Q water and take 0.5-1 μl (0.5-1%) for PCR amplification. There is no need to purify the ligation mixture.
- To a sterile tube, add 10 μls of sterile distilled water; 2 μls of 10X PCR reaction buffer (with MgCl2; add appropriate amount of MgCl2 in case the buffer does not have it); 1.5 μls of 10 mM dNTP mix (180 mM final concentration); 0.5 μls of 10 μM primers (0.25 mM each); 0.5 units of Taq DNA polymerase; 0.5-1 μls of template (diluted ligation mixture) and make up the volume to 20 μls with additional sterile distilled water.
- Mix gently by pippeting and centrifuge briefly to get the contents at the bottom of the tube.
- Place the tube(s) in a PCR machine when the temperature of the block is at about 75°C (manual hot start; this step can be avoided by using a hot start Taq polymerase).
- Carry out PCR according to the following cycling conditions:
- Initial denaturation (1 cycle): 95°C for 1 min.
- 35 cycles of: 95°C for 30 sec, 60°C for 20 sec and extension at 72°C for 1 min.
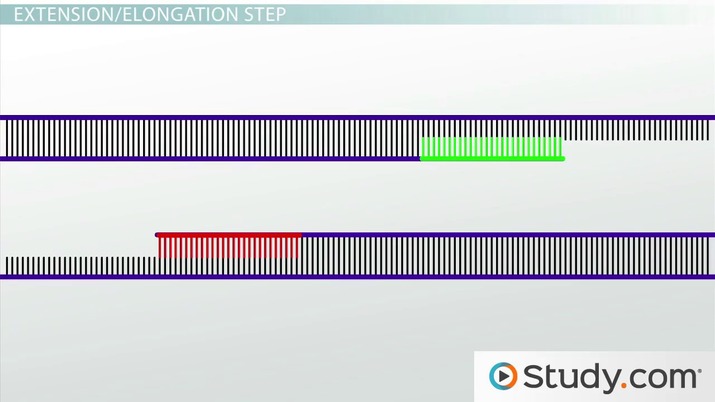
- The PCR products (referred here as Lig-PCR products) can be analyzed immediately by conventional agarose gel electrophoresis or can be stored at -20°C.
- Analysis of Lig-PCR products: Proper controls will have to be used for identifying products derived only from recombinant molecules in the legation mixture(s). Control 1: Circular plasmid, without any insert; Control 2: Only insert(s), without any vector sequence; Control 3: No templates. In cases where the identities of the insert(s) are to be determined, the Lig-PCR products can be directly eluted from the gel, purified and sequenced (preferably cycle sequencing) using either of the primers that were used for PCR.
- Transformation: To confirm the type of recombinant molecules generated in the ligation reaction (and as analyzed using the Lig-PCR method), a small portion of the ligation mixture can be transformed into competent cells and plated on growth media to obtain bacterial colonies. Recombinants can be screened using suitable methods, such as Blue-White screening on X-gal or X-gal/IPTG media for plasmids having the alpha complementation properties of LacZ. The amounts of recombinant and vector derived colonies can be correlated to the band intensities of the corresponding Lig-PCR products.
References
- Lehman IR. DNA ligase: structure, mechanism and function. Science. 1974;186:790–797. [PubMed] [Google Scholar]
- Lindhal T, Wood RD. Quality control by DNA repair. Science. 1999;286:1897–1905. [PubMed] [Google Scholar]
- Maniatis T, Hardison RC, Lacy E, Lauer J, O'Connell C, Quon D, Sim GK, Efstratiadis A. The isolation of structural genes from libraries of eukaryotic DNA. Cell. 1978;15(2):687–701. [PubMed] [Google Scholar]
- Okayama H, Berg P. High-efficiency cloning of full-length cDNA. Mol Cell Biol. 1982;2(2):161–170.[PMC free article] [PubMed] [Google Scholar]
- Hoogenboom HR, Griffiths AD, Johnson KS, Chiswell DJ, Hudson P, Winter G. Multi-subunit proteins on the surface of filamentous phage: methodologies for displaying antibody (Fab) heavy and light chains. Nucleic Acids Res. 1991;19(15):4133–4137.[PMC free article] [PubMed] [Google Scholar]
- Landegren U, Kaiser R, Sanders J, Hood L. A ligase mediated gene detection technique. Science. 1988;241:1077–1080. [PubMed] [Google Scholar]
- Vos P, Hogers R, Bleeker M, Reijans M, Lee T, Hornes M, Frijters A, Pot J, Peleman J, Kuiper M, Zabeau M. AFLP: A new technique for DNA fingerprinting. Nucleic Acid Res. 1995;23:4407–4414.[PMC free article] [PubMed] [Google Scholar]
- Monib Z, Takamor N, Gregg N, Timothy M, Philip P, Francis B. Ligase based detection of mononucleotide repeat sequences. Nucleic Acid Res. 1999;27:e4–e4.[Google Scholar]
- Beaudry D, Perreault JP. An efficient strategy for the synthesis of circular RNA molecules. Nucleic Acids Res. 1995;23:3064–3066.[PMC free article] [PubMed] [Google Scholar]
- Drumm ML. Chromosome jumping and linking libraries in E.coli. In: Dracopoli NC, Hainer JL, Korf BR, Morton CC, Seidman CE, Seidman JG, editors. Current Protocols in Human Genetics. New York: John Wiley & Sons; 1994. Unit 5.4.
- Pulido JC, Duyk GM. Construction of small insert libraries enriched for short tandem repeat sequences by marker selection. In: Dracopoli NC, Hainer JL, Korf BR, Morton CC, Seidman CE, Seidman JG, editors. Current Protocols in Human Genetics. New York: John Wiley & Sons; 1994. Unit 2.2.
- Klickstein LB. Production of substracted cDNA library. In: Ausubel FM, Brent R, Kingston RE, Moore DD, Seidman JG, Smith JA, Struhl K, editors. Current Protocols in Molecular Biology. New York: John Wiley & Sons; 1988. Unit 25B.1.
- Odell M, Shuman S. Footprinting of chlorella virus DNA ligase bound at a nick in duplex DNA. J Biol Chem. 1999;274:14032–14039. doi: 10.1074/jbc.274.20.14032. [PubMed] [CrossRef] [Google Scholar]
- Doherty AJ, Dafforn TR. Nick recognition by DNA ligase. J Mol Biol. 2000;296:43–56. doi: 10.1006/jmbi.1999.3423. [PubMed] [CrossRef] [Google Scholar]
- Courtney BC, Williams KC, Bing QA, Schlager JJ. Capillary gel electrophoresis as a method to determine ligation efficiency. Anal Biochem. 1995;228:281–286. doi: 10.1006/abio.1995.1351. [PubMed] [CrossRef] [Google Scholar]
- Tang Z, Wang K, Tan W, Li J, Liu LF, Guo Q, Meng X, Ma C, Huang S. Real-time monitoring of nucleic acid ligation in homogenous solutions using molecular beacons. Nucleic Acids Res. 2003;31(23):e148–e148. doi: 10.1093/nar/gng146.[PMC free article] [PubMed] [CrossRef] [Google Scholar]
- Jansson V, Jansson K. An enzymatic bioluminescent assay for DNA ligation. Analytical Biochem. 2004;324:307–308. [PubMed] [Google Scholar]
- Mullis K, Faloona F, Scharf S, Saki R, Horn G, Erlich H. Specific enzymatic amplification of DNA in vitro: the polymerase chain reaction. Biotechnology. 1986;24:17–27. [PubMed] [Google Scholar]
- Messing J. New M13 vectors for cloning. Methods Enzymol. 1983;101:20–78. [PubMed] [Google Scholar]
- Sambrook J, Fritsch EF, Maniatis T. Molecular Cloning: A Laboratory Manual. New York: Cold Spring Harbor Laboratory Press; 1989. Vol. 1, 2, 3.
Articles from Biological Procedures Online are provided here courtesy of BioMed Central
